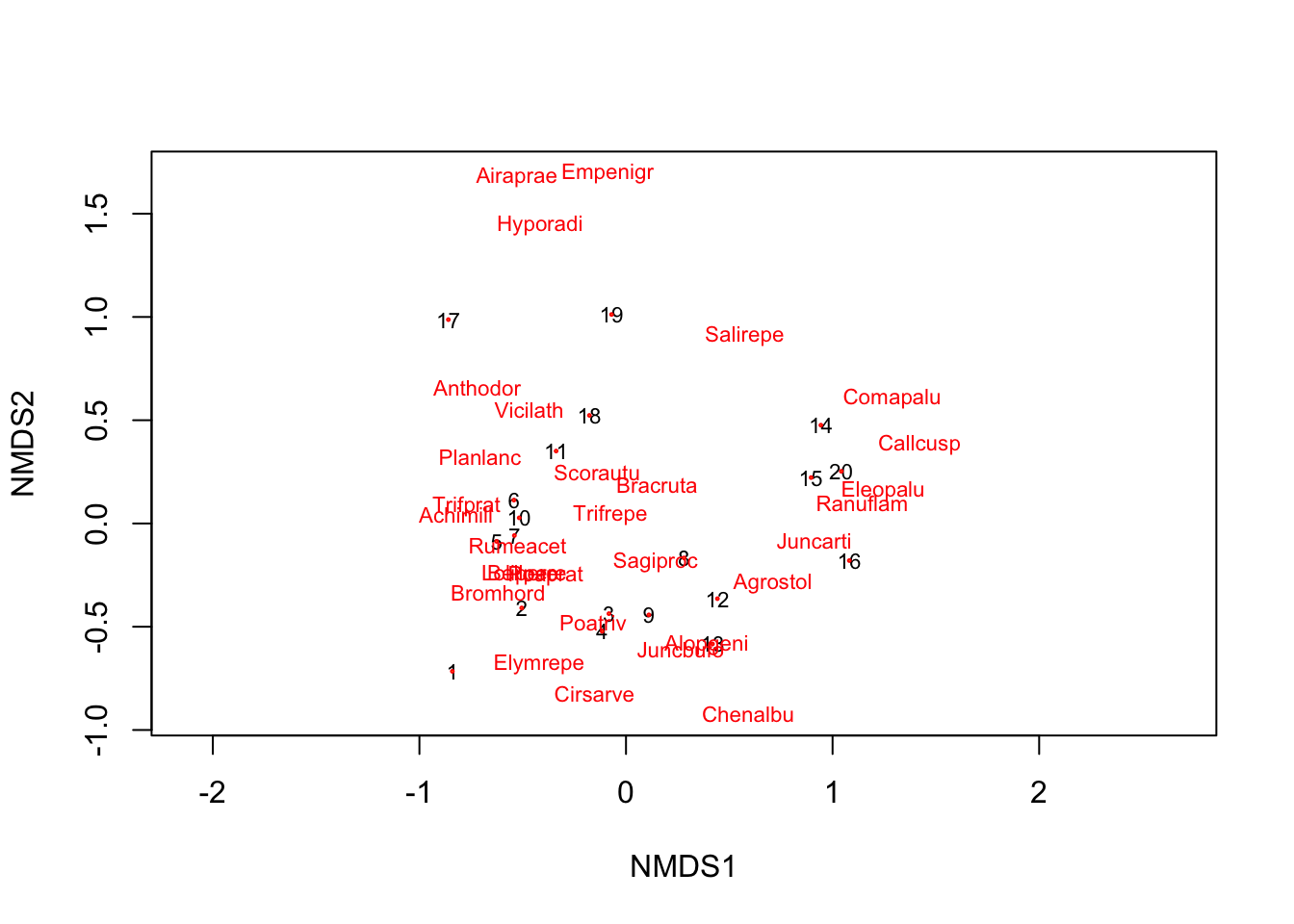
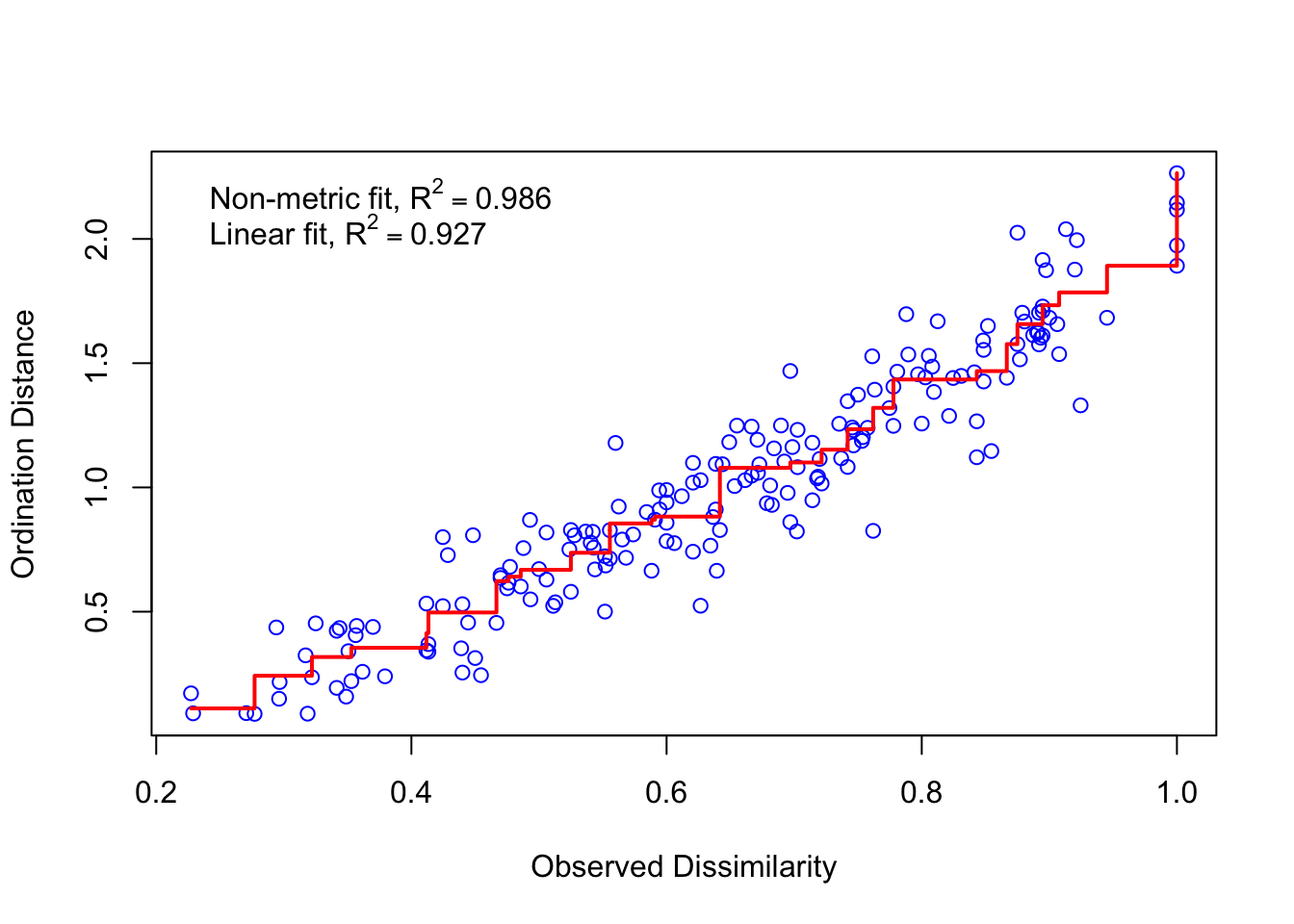
Rows: 20
Columns: 30
$ Achimill <dbl> 1, 3, 0, 0, 2, 2, 2, 0, 0, 4, 0, 0, 0, 0, 0, 0, 2, 0, 0, 0
$ Agrostol <dbl> 0, 0, 4, 8, 0, 0, 0, 4, 3, 0, 0, 4, 5, 4, 4, 7, 0, 0, 0, 5
$ Airaprae <dbl> 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 2, 0, 3, 0
$ Alopgeni <dbl> 0, 2, 7, 2, 0, 0, 0, 5, 3, 0, 0, 8, 5, 0, 0, 4, 0, 0, 0, 0
$ Anthodor <dbl> 0, 0, 0, 0, 4, 3, 2, 0, 0, 4, 0, 0, 0, 0, 0, 0, 4, 0, 4, 0
$ Bellpere <dbl> 0, 3, 2, 2, 2, 0, 0, 0, 0, 2, 0, 0, 0, 0, 0, 0, 0, 2, 0, 0
$ Bromhord <dbl> 0, 4, 0, 3, 2, 0, 2, 0, 0, 4, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0
$ Chenalbu <dbl> 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0
$ Cirsarve <dbl> 0, 0, 0, 2, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0
$ Comapalu <dbl> 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 2, 2, 0, 0, 0, 0, 0
$ Eleopalu <dbl> 0, 0, 0, 0, 0, 0, 0, 4, 0, 0, 0, 0, 0, 4, 5, 8, 0, 0, 0, 4
$ Elymrepe <dbl> 4, 4, 4, 4, 4, 0, 0, 0, 6, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0
$ Empenigr <dbl> 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 2, 0
$ Hyporadi <dbl> 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 2, 0, 0, 0, 0, 0, 2, 0, 5, 0
$ Juncarti <dbl> 0, 0, 0, 0, 0, 0, 0, 4, 4, 0, 0, 0, 0, 0, 3, 3, 0, 0, 0, 4
$ Juncbufo <dbl> 0, 0, 0, 0, 0, 0, 2, 0, 4, 0, 0, 4, 3, 0, 0, 0, 0, 0, 0, 0
$ Lolipere <dbl> 7, 5, 6, 5, 2, 6, 6, 4, 2, 6, 7, 0, 0, 0, 0, 0, 0, 2, 0, 0
$ Planlanc <dbl> 0, 0, 0, 0, 5, 5, 5, 0, 0, 3, 3, 0, 0, 0, 0, 0, 2, 3, 0, 0
$ Poaprat <dbl> 4, 4, 5, 4, 2, 3, 4, 4, 4, 4, 4, 0, 2, 0, 0, 0, 1, 3, 0, 0
$ Poatriv <dbl> 2, 7, 6, 5, 6, 4, 5, 4, 5, 4, 0, 4, 9, 0, 0, 2, 0, 0, 0, 0
$ Ranuflam <dbl> 0, 0, 0, 0, 0, 0, 0, 2, 0, 0, 0, 0, 2, 2, 2, 2, 0, 0, 0, 4
$ Rumeacet <dbl> 0, 0, 0, 0, 5, 6, 3, 0, 2, 0, 0, 2, 0, 0, 0, 0, 0, 0, 0, 0
$ Sagiproc <dbl> 0, 0, 0, 5, 0, 0, 0, 2, 2, 0, 2, 4, 2, 0, 0, 0, 0, 0, 3, 0
$ Salirepe <dbl> 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 3, 3, 5
$ Scorautu <dbl> 0, 5, 2, 2, 3, 3, 3, 3, 2, 3, 5, 2, 2, 2, 2, 0, 2, 5, 6, 2
$ Trifprat <dbl> 0, 0, 0, 0, 2, 5, 2, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0
$ Trifrepe <dbl> 0, 5, 2, 1, 2, 5, 2, 2, 3, 6, 3, 3, 2, 6, 1, 0, 0, 2, 2, 0
$ Vicilath <dbl> 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 2, 0, 0, 0, 0, 0, 0, 1, 0, 0
$ Bracruta <dbl> 0, 0, 2, 2, 2, 6, 2, 2, 2, 2, 4, 4, 0, 0, 4, 4, 0, 6, 3, 4
$ Callcusp <dbl> 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 4, 0, 3, 0, 0, 0, 3